Genomic info from crossbreds is routinely generated for genomic evaluations. The goal of this research is to analyze autozygosity and genetic differentiation in Landrace by Large-White breeds by utilizing the genotypic info of SNP arrays in 1,173 crossbreds.
A most probability method was developed to estimate the chance of autozygosity (FL ). Regions of differentiation between breeds had been investigated utilizing FST and the distinction in allele frequencies between the two parental breeds (릌Δ) at every single-nucleotide polymorphism (SNP) place.
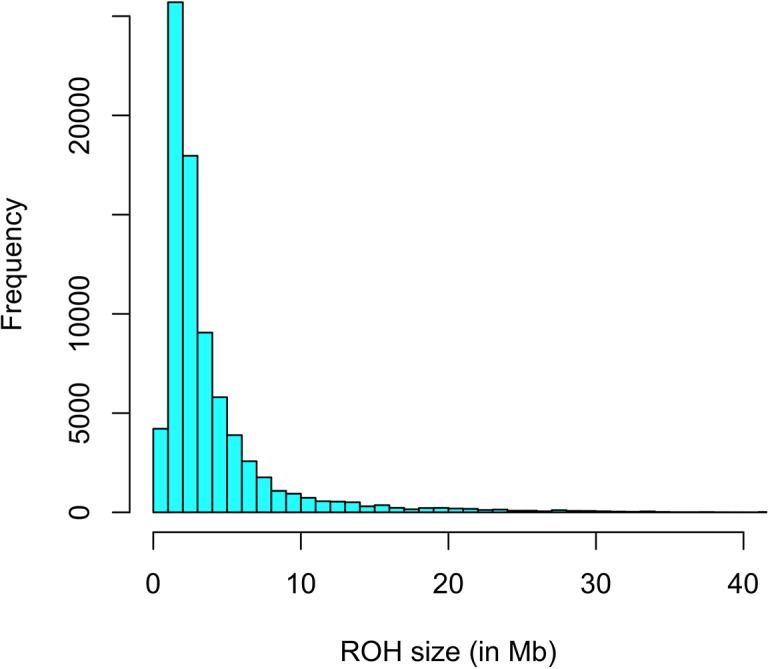
A most probability method was proposed to estimate allele frequencies in the parental populations. The common size of runs of homozygosity (ROH) throughout the genome was 3.91, 2.3, and 0.7 Mb for segments with not less than 25, 15, and 5 SNPs, respectively. Average age to coalesce was 46, 414, and 388 years for segments with not less than 25, 15, and 5 SNPs, respectively.
The chance of autozygosity was not uniform alongside the crossbred genome, being increased at the heart for many chromosomes. The correlation between autozygosity and distance to the closest telomere was constructive and important in most chromosomes, which could possibly be attributed to the increased recombination fee close to telomeres.
We additionally report a comparatively excessive unfavorable correlation between chance of recombination (from a printed map) and chance of autozygosity. It helps that structural traits of the chromosomes associated to recombination fee decide autozygosity at every chromosomal place of the pig genome.
The common is Δ throughout the genome was 0.17 (SD = 0.16). After testing for variations in allele frequencies between the parental breeds, there have been 4,184 SNPs with a probability ratio check, LRT ≥ 32.02. The common FST throughout the genome was 0.038 (SD = 0.059).
There had been 2,949 SNPs with FST > 0.125. The correlation between estimates of FL and estimates of FST throughout the genome was -0.10 (SE = 0.006). Analysis of the gene content material of the genomic areas with the 2000 SNPs with highest LRT for FL and excessive FST confirmed overrepresentation of genes with a regulatory perform.
Genes with organic capabilities related to manufacturing, such as tissue improvement, anatomical construction, and animal organ improvement, had been additionally overrepresented in areas with a excessive FST

Relationships between estimated autozygosity and complicated traits in the UK Biobank.
Inbreeding will increase the threat of sure Mendelian problems in people however can also cut back health by means of its results on complicated traits and ailments. Such inbreeding melancholy is assumed to happen resulting from elevated homozygosity at causal variants which might be recessive with respect to health.
Until just lately it has been tough to amass giant sufficient pattern sizes to analyze the results of inbreeding melancholy on complicated traits utilizing genome-wide single nucleotide polymorphism (SNP) information in population-based samples. Further, it’s tough to deduce causation in analyses that relate diploma of inbreeding to complicated traits as a result of confounding variables (e.g., schooling) might affect each the probability for fogeys to outbreed and offspring trait values.
The current research used runs of homozygosity in genome-wide SNP information in as much as 400,000 people in the UK Biobank to estimate the proportion of the autosome that exists in autozygous tracts-stretches of the genome that are equivalent resulting from a shared widespread ancestor.
After a number of testing corrections and controlling for attainable sociodemographic confounders, we discovered important relationships in the predicted course between estimated autozygosity and three of the 26 traits we investigated: age at first sexual activity, fluid intelligence, and pressured expiratory quantity in 1 second. Our findings corroborate these of a number of printed research.
These outcomes might suggest that these traits have been related to Darwinian health over evolutionary time. However, some of the autozygosity-trait relationships had been attenuated after controlling for background sociodemographic traits, suggesting that different explanations for these associations haven’t been eradicated.
Care must be taken in the design and interpretation of ROH research so as to glean dependable details about the genetic structure and evolutionary historical past of complicated traits.
